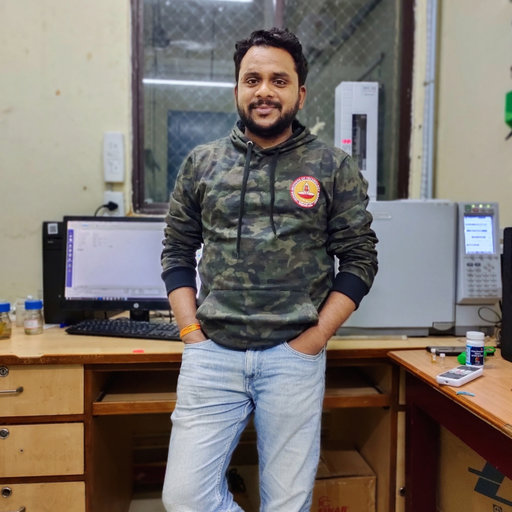

- Mestrenova stacked spectra how to#
- Mestrenova stacked spectra manual#
- Mestrenova stacked spectra verification#
- Mestrenova stacked spectra license#
Baseline correction should be applied AFTER phasing.Watch the peaks at the other end of the spectrum away from the pivot while adjusting 1st-order phase. When to apply a baseline correction in 2D NMR? t1 and t2 refer to time-domain data f1 and f2 refer to frequency domain data after FT of t1 and t2 dimensions. What are the dimensions of a 2D NMR spectrum?ĢD NMR Spectrum Processing with Mnova The two dimensions in a 2D spectrum is specified as t2or f2(horizontal) and t1or f1(vertical) dimensions, respectively. Check & Select appropriate window functions for t2 and t1 under Processing->Apodization 2D spectrum should be processed in the following order in Mnova: 1. Select either f2or f1from the top menu buttons before applying the processing command. The processing command below is often applied to f2 and f1 (or t2 and t1) separately. To symmetrize, use the menu combination: Processing > Symmetrize > COSY-like.For symmetrize to be possible, both dimensions need to have the same digital resolution (same number of points), so you may need to zero-fill the t1/f1dimension up to the same. Mnova provides an interface for processing and plotting NMR data. MNova: 2D Spectrum Processing page 3 For a magnitude mode COSY and related symmetrical spectra, you can symmetrize, to remove t1 noise. The predicted spectrum is stacked with the experimental one for visual comparison. Processing data with Mestrelab Mnova This exercise has three parts: a 1D 1H spectrum to baseline correct, integrate, peak-pick, and plot a 2D spectrum to plot with a 1H spectrum as a projection and three 1H spectra to compare and plot against the same axes. Overview of Mestrelab and Mnova Open and process 1D and 2D NMR data.
Mestrenova stacked spectra how to#
How to process NMR data with Mestrelab Mnova?
Mestrenova stacked spectra license#
Standard academic license price for academia for Mnova Suite is aprox. Mnova for Undergraduate Schools represents a huge saving. The GUI that pops up allows you to annotate the peak with any text. The shortcut for this mode is ‘Shift + P’. Click ‘grid’ and then uncheck all of the boxes. On the ‘Scales’ tab, unclick vertical on the axes section.
Right click on the spectra and click ‘Properties’. Remove all the grid lines and vertical scales from the spectra. Most commands that are executed depend on the values of certain parameters. For example, use File / Open to open a dataset, or type efp on the command line to process a 1D dataset. TopSpin commands can be input either by typing them on the command line or by using the Flowbars to select the command. It is responsible for the often complex splitting of resonance lines in the NMR spectra of fairly simple molecules. Most importantly, J-coupling provides information on the connectivity of chemical bonds. In NMR spectroscopy, J-coupling contains information about relative bond distances and angles. integration, peak analysis) using Varian Topspin 3 and Mestrenova. In pure D 2 O, exchangeable protons are completely replaced by deuterium, effectively removing the signal for those protons from the NMR spectrum by reducing its intensity to near zero.
Mestrenova stacked spectra verification#
Mnova NMR is a basic plugin containing the advanced functionality offered by the advanced plugins available within Mnova such as mixtures analysis, reaction monitoring, quantitation, chemical shift prediction, screening, verification as well as physico-chemical properties prediction. 13C NMR spectra are referenced to the residual solvent peak for DMSO-d6 (1H: 1.94 ppm. It is also possible to easily change the color of the spectrum background to suit your eyesight, your working hours, or your mood. Just select the desired spectra on the page navigator (by holding down ‘CTRL key’ while clicking on each spectrum) and then issue the command ‘Stack/Superimpose spectra’. page 25 D.How do you stack NMR spectra on MestreNova? page C, HSQC, and other x-nuclei Prediction. Since the claimed emission lines lie around 3. Crosshair Tool page NMR Prediction - 1 H. matter, motivated by recent claims of unidenti ed emission lines in the stacked X-ray spectra of galaxy clusters and the centers of the Milky Way and M31. New Spectrum (without Solvent or Impurity Peaks) page Coupling Constants. page Split Partially Overlapping Multiplets page 18 A. page 10 4) Report Multiplets for journals, papers, patents.
Mestrenova stacked spectra manual#
page 7 1) Automatic Integration page 8 2) Manual Integration. Peak Picking page 6 1) Automatic Peak Picking page 6 2) Manual Theshold. 1 MestReNova Guide Center for Advanced Materials University of Massachusetts Lowell Training and Operation Manual By Wendy Gavin NMR Technician 3/2014 1Ģ 1.
